- Home
Welcome !
Welcome to my home page for this particular lecture series. This site is designed as a hub for curating and sharing of lectures and lecture material, course syllabi, assignments and links to relevant resources. The web-site is maintained by me. Therefore, material on this site normally supersedes all other course information found elsewhere (such as icollege). You can use the menu bar that is located below the course logo to navigate through the site and to find potential links that may be pertinent to this course -or research and study in the Biological Sciences in general.

- Courses
- Resources
General Resources

Mechanisms of Cellular Response to Environmental Changes
|
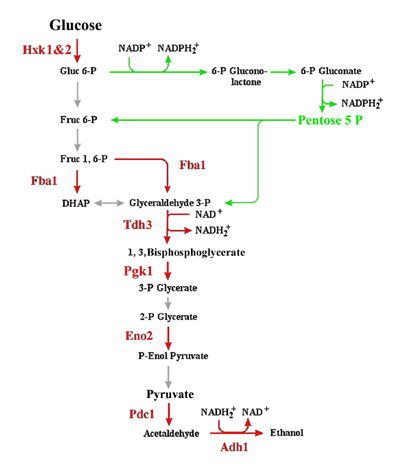
Research in the laboratory is centred around mechanisms that underpin cellular response(s) to changes in the environment and phases of growth or life cycle. The research has two quite distinct components: the one uses Saccharomyces cerevisiae as a model eukaryotic system to evaluate the cellular response to heavy metal exposure, such as Copper, Chromium (VI) and Cadmium. The other, in collaboration with Professor P.C.Tai (also in the Dept of Biology at GSU) investigates the various roles of SecA in protein tranlocation in bacteria. Oxidative Targeting of Enzymes in S. cerevisiae: While both cellular response mechanisms differ dramatically in their mechanisms of action, they each share a significant metabolic component. The first level response of S. cerevisae to exposure to sub-lethal concentrations of heavy metals involves targeted oxidation of enzymes predominantly within the glycolytic pathway (see Figure 1). |
|
|
||
| Figure 1: The Glycolytic and Pentose Phosphate pathways in S. cerevisiae, showing the key enzymes and reactions (in red) that appear to be specifically targeted for oxidation in response to the presence of heavy metals, such as Cu, Cd and Cr. | ||
|
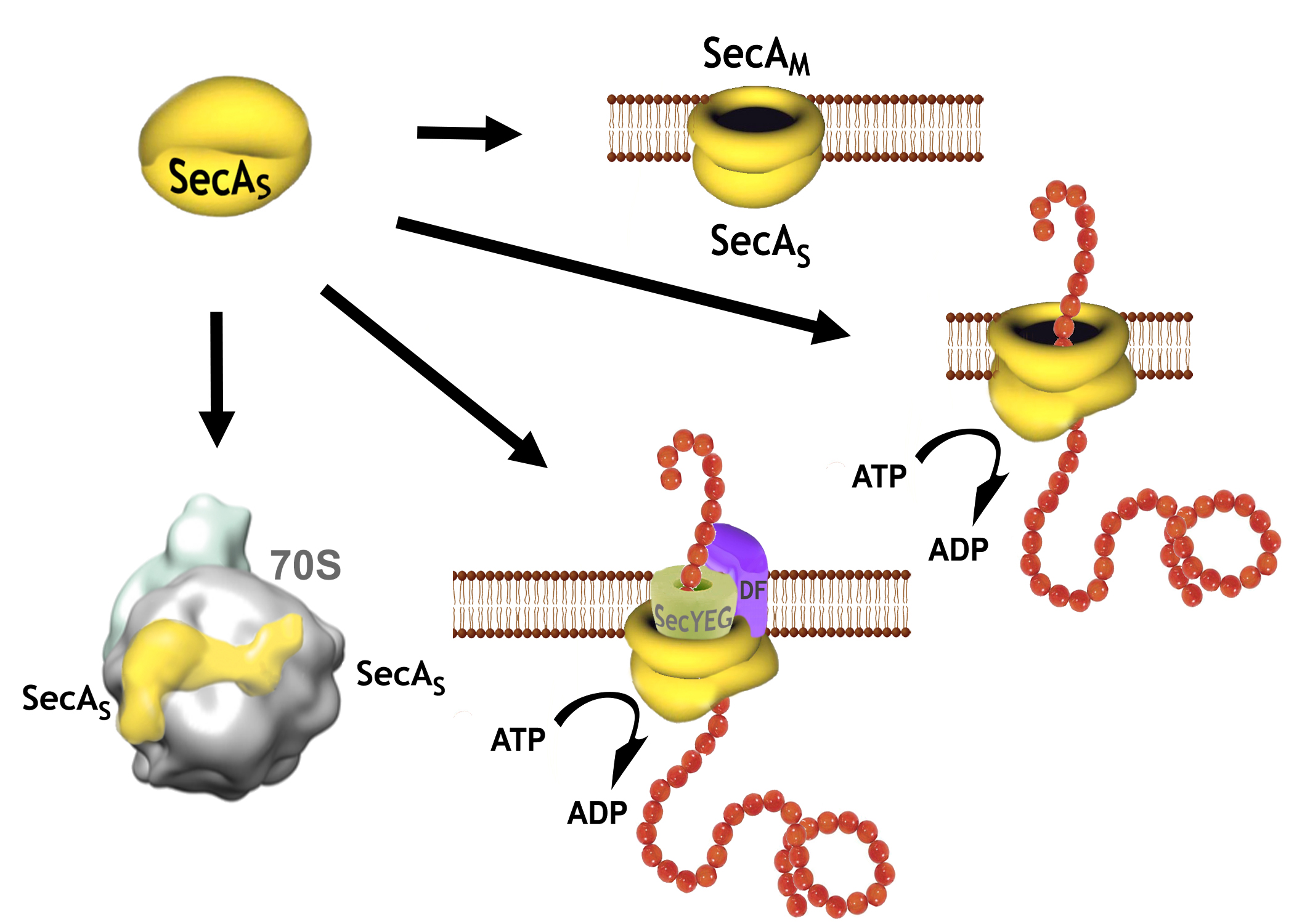
Roles of the multifunctional SecA protein in protein translocation: SecA is a key component of the translocation machinery in bacteria, and while it has s no direct analogue in eukaryotes provides a number of essential roles in protein translocation in the cell; from a requisite ATPase in the SecA-YEG transporter complex that drives protein precursors/peptide through the SecYEG protein translocation channels to forming an additional "SecA-only" channel that facilitates the translocation of signal free proteins across the cytoplasmic membrane -albeit with less efficiency than the SecA-YEG channels (Figure 2). Expression of SecA protein is also distingct from theat of all the other members of the translocation machinery, and has been found to be differentially expressed at various phase of cellular growth in E. coli and Bacillus sp.. As such, it is believd that the role of SecA may vary as a functon of growth phase. An analysis of this role for SecA forms the basis of reseach in this laboratory, in strong collaboration with P.C. Tai, who has been a pioneer in this area of research for the last forty years and more.. |
||
|
Figure 2. Some of the proposed multi-protein complexes that SecA is believed to form with itself -as a soluble SecAs dimer or in the membrane as SecA-only channel, with SecYEG,DF and with the 70S ribosome. |
|
|
| .
|
||
|
|
||