- Home
Welcome !
Welcome to john houghton's home page for his biology courses. This site is designed as a hub for curating and sharing lectures, course syllabi, assignments, and links to relevant resources. Use the menu bar at the top of the screen to navigate through the site.
(Please note: this page is currently under construction.)

- BIOL 2107
Fall '23 CRN86772
Lectures: (1)
- Courses
BIOL 2107 Principles of Biology I
- Resources
General Resources
It must be able to store all of an organism's genetic information.It must be susceptible to mutation.It must be precisely replicated in the cell division cycle.It must be expressible as the phenotype.
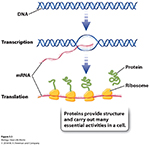
There are many steps between genotype and phenotype; as genes cannot (by themselves) directly produce a phenotype, with the first Porcess involvling the synthesis of RNA.
Review how does RNA differ from DNA
RNA is usually single-stranded. The sugar in RNA is ribose,not deoxyribose.
RNA can fold over and base-pair with itself.
RNA comes in various forms/sequences, commensurate with function.
Synthesis of RNA:----- Transcription is a "DNA-Directed"process.
It is carried out by RNA polymerase, which is an enzyme that uses DNA as a template to make RNA.
As the RNA transcript forms, it peels away, allowing the already transcribed DNA to be rewound into the double helix.
Initiation of transcription requires a "promoter" and an RNA polymerase.
There is at least one "promoter" for each gene that is to be transcribed into mRNA.
The RNA polymerase binds to the promoter region when conditions allow.
The promoter sequence directs the RNA polymerase as to which of the double strands is the template and in what direction the RNA polymerase should move.
In effect, the promoters serve as "punctuation marks" for the transcriptional process.
As in replication of DNA, transcription of RNA is always synthesized in the 5' to 3' direction
Not all promoters are identical. Some bind RNA polymerase more effectively than others; which causes them to be transcribed more frequently........when conditions allow.
Prokaryotes have only ONE type of RNA polymerase which transcribes ALL THREE types of RNA, mRNA, tRNA, and rRNA.
Eukaryotes have THREE DIFFERENT RNA polymerases: RNA polymerase I, II, and III.
It is the RNA polymerase II that makes ALL the mRNA in eukaryotes.
In eukaryotic cells, other proteins must also bind to the DNA around the promoter to prepare a 'docking site' for the transcribing enzyme to bind. This uniquely eukaryotic feature provides a way to regulate the transcription of particular genes in a uniquely eukaryotic way.
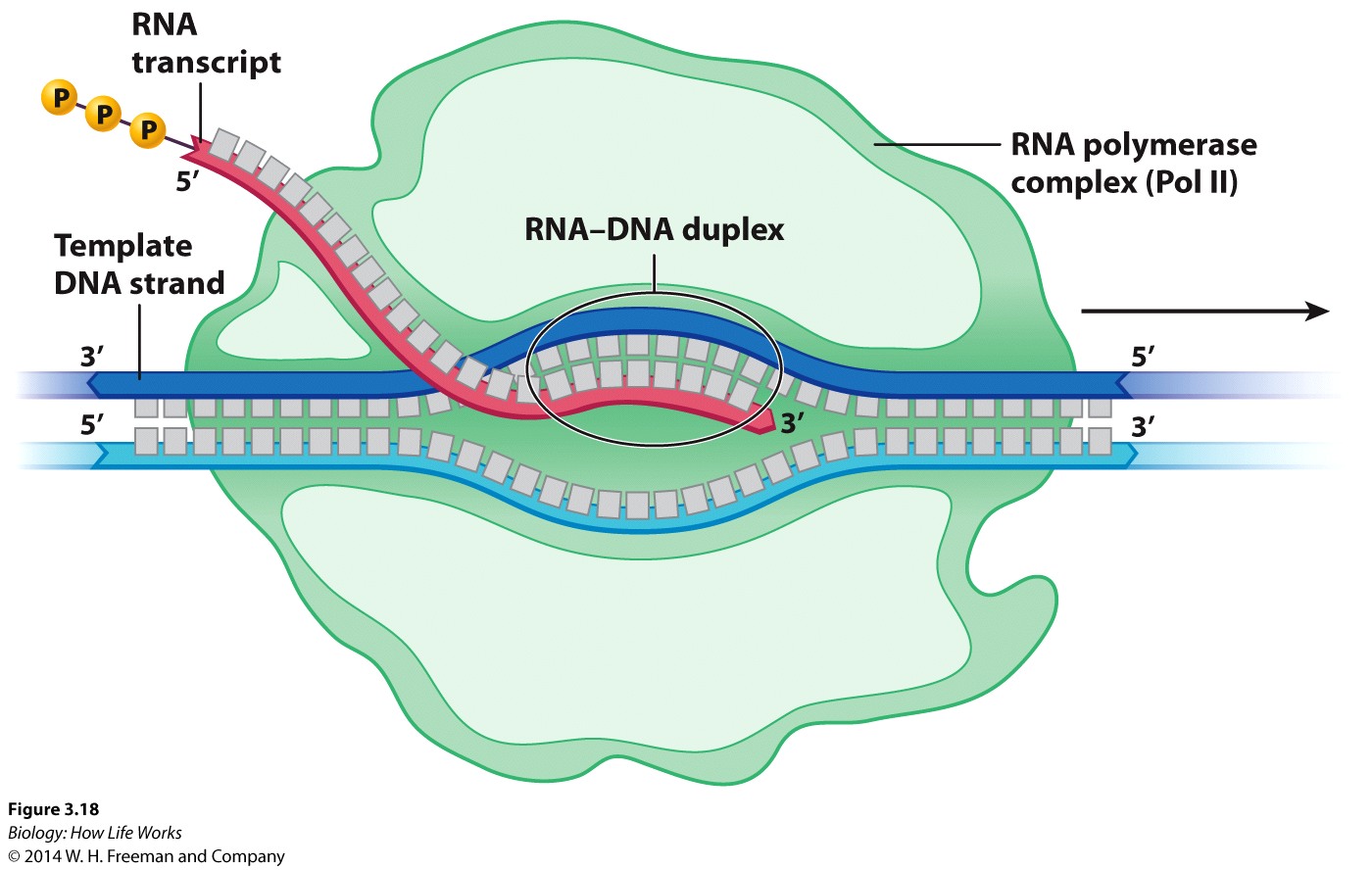
RNA polymerase elongates the transcript.
After binding, RNA polymerase unwinds the DNA about 20 base pairs at a time and READS the template strand in the 3' to 5' direction
The RNA transcript is thus extended in an anti-parallel manner with respect to the DNA strand that acts as the "template" strand.
Energy for synthesis comes from the removal and breakdown of the pyrophosphate group from each nucleotide added
Transcription errors for RNA polymerases are much higher (relative to DNA polymerases);making a mistake every 104 to 105 bases incorporated into the growing RNA strand.
Transcription terminates at particular base sequences in the DNA that specify termination.
RNA Transcription, and Gene expression in Eukaryotes is far more complex than it is in Prokaryotes
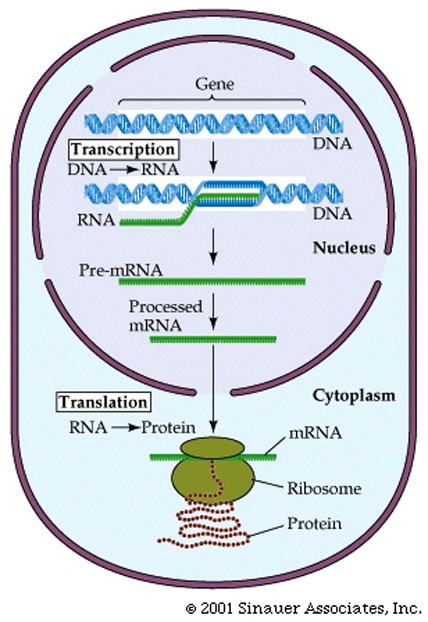
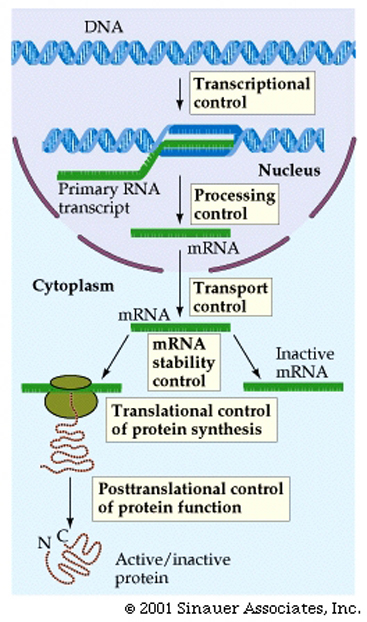
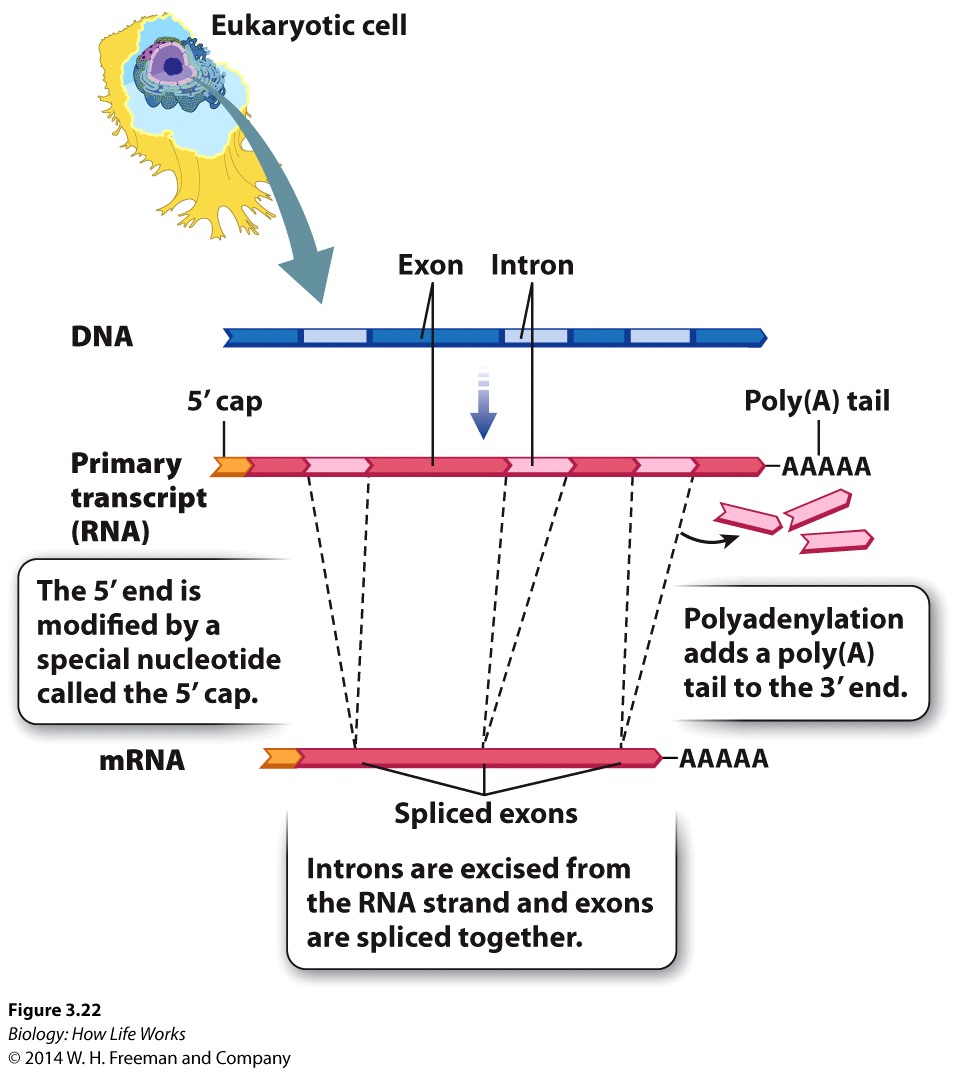
The mRNA that gives rise to the protein product in prokaryotes is said to be co-linear withthe DNA sequence from which it came.
This theme varies somewhat for eukaryotes.
1. 5' end of the mRNA is "capped" or modified to establish it as being the 5' end of a mRNA, eventually gives rise to the mature mRNA
2. Distinct lack of colinearity of mRNA (ie the final RNA product is NOT the same length as the initial mRNA that is synthesized from the DNA) and is usually much smaller than the originally syntheisized hnRNA.
3. Very often mature mRNA sequences (or pre-RNA sequences) encode for a “3' untranslated region” (UTR)that (as the name suggests) is transcribed, BUT NOT translated, and is very often "tagged" by a series of "AAAAAAA" repeats. or a "poly A Tail" to round out the distinctlt eukaryotic maturation process
Consequently many of the major differences between Proks and Euks are defined by the separation of translation from the other aspects of the central dogma by the nuclear membrane.
The GENETIC CODE
DNA codes for RNA through the transcriptional process.
mRNA is a single stranded nucelic acid that is converted into a polypeptide chain through a "3 for one" genetic code. The mRNA is "read" in three-base, contiguous segments called codons (equivalent to "words").
The number of different codons possible is 64, because each position in the codon can be occupied by one of four different bases.

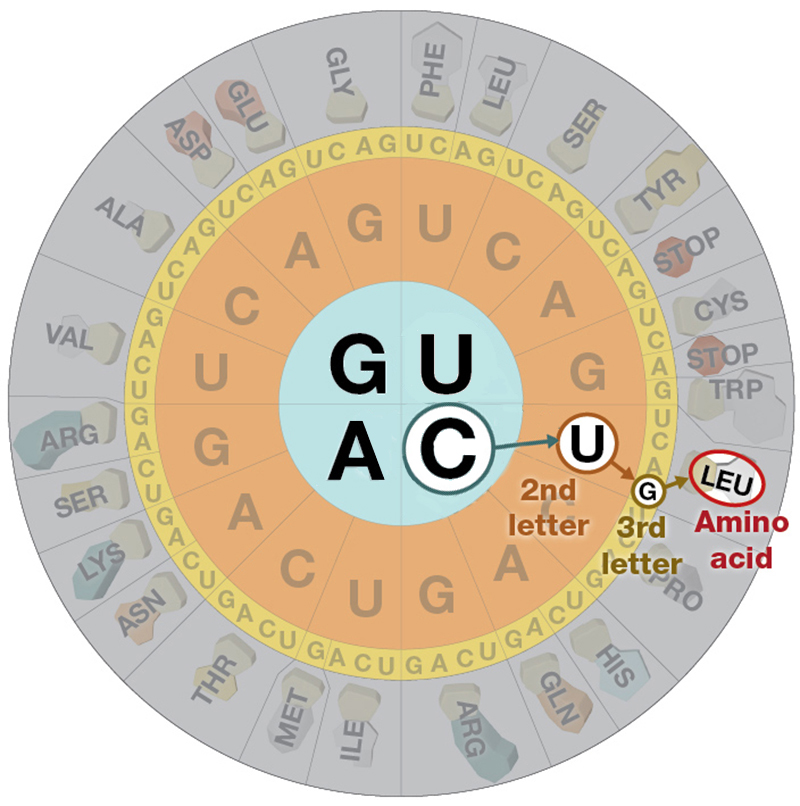
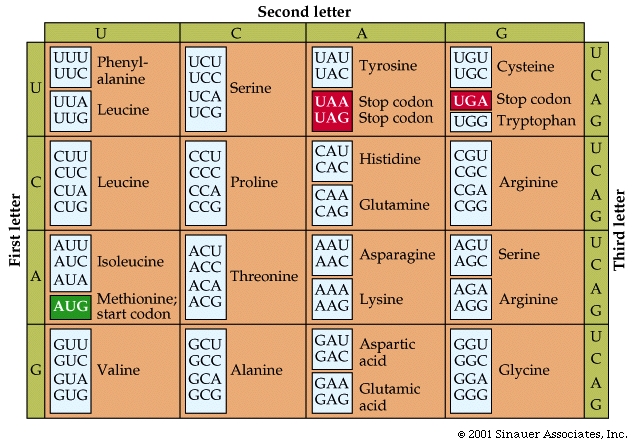

Four possibilities for the first base, times four for the second, times four for the third yields 64 possibilities.
In the genetic code, there are no commas or punctuation marks,it is therefore very important to know where to BEGIN and where to END the process. For this reason there are always well defined START and STOP "codons".
L / ETT / HEC / ATG / ETT / HER / ATO / FFT / HEB / AT
LE / TTH / ECA / TGE / TTH / ERA/ TOF / FTH / EBA / T
LET / THE / CAT / GET / THE / RAT / OFF / THE / BAT
The 64 possible codons (or "words") code for only 20 amino acids as well as the start and stop signals found in ALL mRNA molecules.
AUG, which codes for the amino acid methionine, is called the start codon, which initiates the translational process.
Three of the possible codons are stop< codons (UAA, UAG, and UGA), which direct the ribosomes to STOP reading the mRNA; that is, they end translation.
The
genetic code is redundant but not ambiguous. This means that many amino acids have more than one codon, but only one amino acid is specified for any one codon.
The genetic code is nearly Universal, applying to all species on our planet. (which would argue strongly for an evolution. Why??)
Minor variations are found, however, within mitochondria and chloroplasts; but all other exceptions are few and far between.
Translation occurs at ribosomes, which are molecular protein synthesizing machines that hold mRNA and tRNA in place.
Translation of the RNA into polypeptides.
In prokaryotes, translation of the mRNA often begins before transcription is complete, as we will discuss in the next (and "last") lecture.
In eukaryotes, however, because the whole process is far more complicated and Replication (together with Transcription and maturation of the transcript) is separated from Translation in the cytoplasm or "Edoplasmic Reticulum") by the nucelar membrane... translation can olny begin in Eukaryotes AFTER transcription and maturation of the RNA is complete. Thereafter, however, translation is very similar in both Eukaryotes and Prokaryotes, except perhaps) that in Eukaryotes the Ribosome complexes are a little bit larger than they are in Prokaryotes.
But let's start with the basics.
Translation occurs at ribosomes, which are molecular protein synthesizing machines that hold mRNA and tRNA in place.
To ensure protein specificity there is a very faithful "activation" of all the tRNA's in the cell:
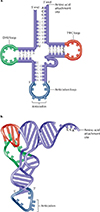
The tRNA's must read mRNA correctly.The tRNA's must carry the correct amino acids.Transfer RNA's carry specific amino acids and bindto specific codonsSpecific tRNA molecules function as adapters.Each carries an amino acid, associated with mRNA molecules, and interacts with ribosomes.A tRNA molecule has 75 to 80 nucleotides.At the 3' end of every tRNA molecule is a site to which its specific amino acid binds covalently.
At about the midpoint in the molecule are three bases called the anticodon. This anticodon is the "contact point" between the tRNA and the mRNA, and is complementary (AND anti parallel) to the mRNA codon.
The codon and anticodon unite by complementary base pairing.

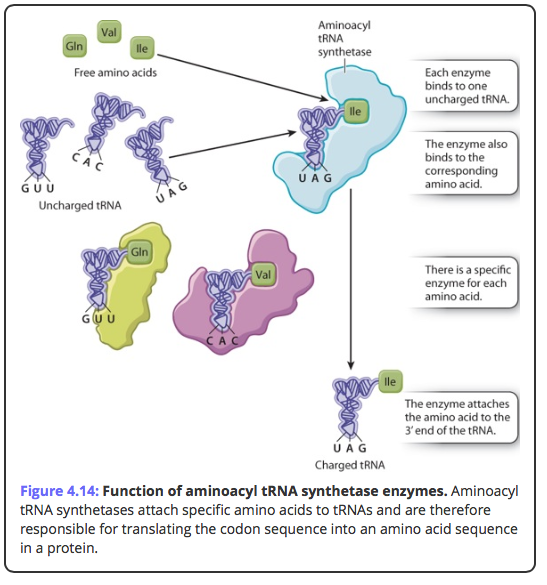
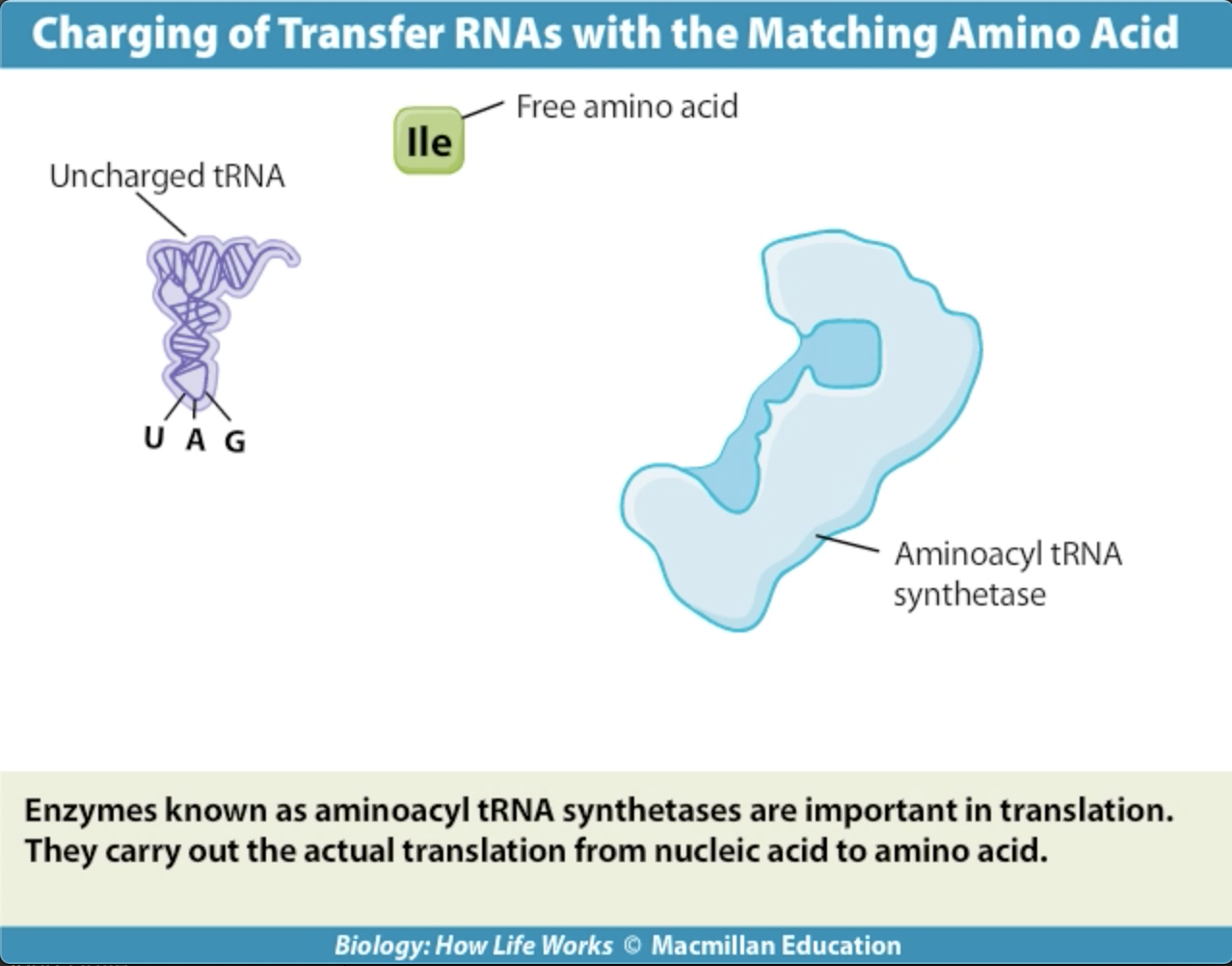
Activating enzymes link the appropriate tRNA's and thei corresponding amino acids.
The correct amino acids are attached to the correct tRNA's by a family of activating enzymes called aminoacyl-tRNA synthetases. Each activating enzyme is specific for one amino acid and its tRNA.The enzyme has a three-part active site that binds a specific amino acid, ATP, and a specific tRNA, which is charged with a high-energy bond. The reactions have two steps:
Enzyme + ATP + amino acid --> enzyme-AMP-aa + PPiEnzyme-AMP-aa + tRNA --> enzyme + AMP + tRNA-aa
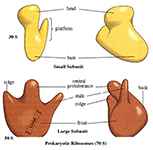
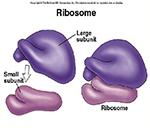
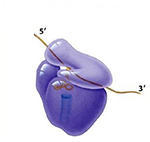
Each ribosome has two subunits, a larger one and a smaller one.
The large one in eukaryotes has three different associated rRNA molecules and 45 different proteins.
The smaller subunit has one rRNA and 33 different protein molecules.
When they are not translating, the two subunits are separate.
Ribosomes are simply molecular factories and are nonspecific. They combine with any mRNA and all tRNA's.
The large subunit has effectivley three sites where tRNA molecules bind.
The A site is where the tRNA anticodon binds to the mRNA codon.
The P site is where the tRNA adds its amino acid to the growing polypeptide chain.
The E (exit) site is where the tRNA, less its amino acid, goes before leaving the ribosome
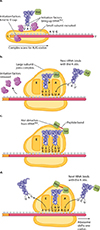
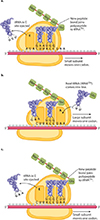
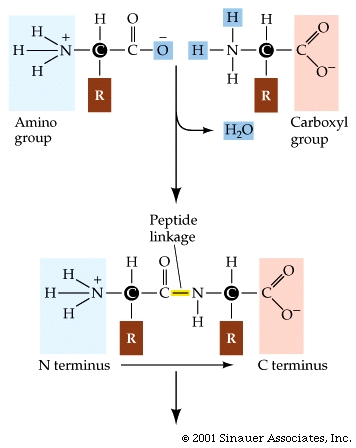
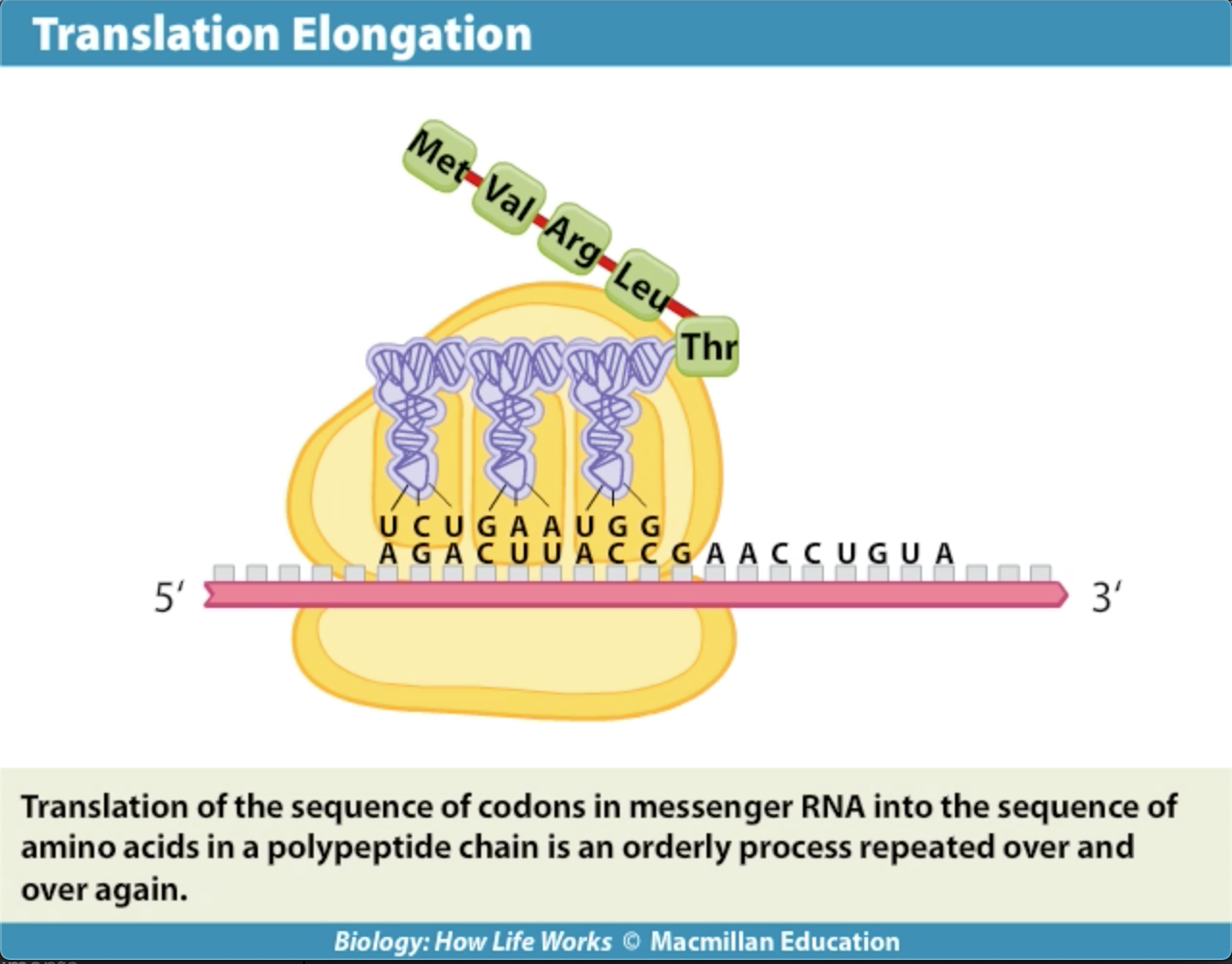
How does the procees get started....
Translation of mRNA often involves multiple rounds of translation, which can occur simultaneously, with mRNA that is being translated often looking like "beads on a string".
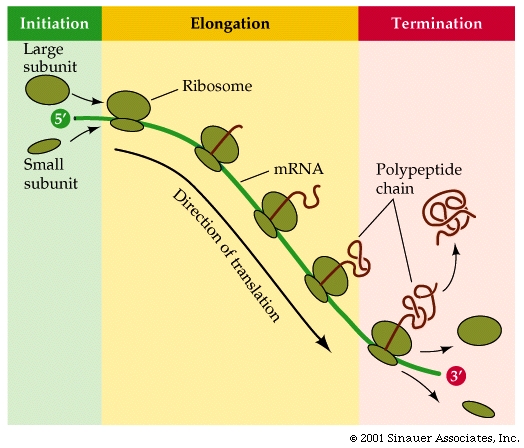
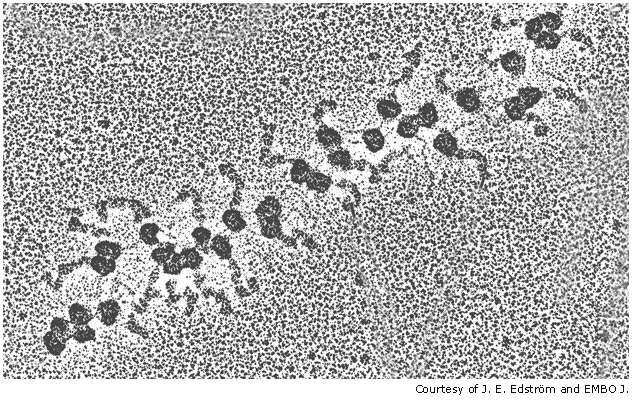
Once translation has been completed, the newly completed polypeptide then separates from the ribosome, folds in to its apporpriate protein configuration and is directed to the part of the cell where it is best put to use to undertake its function.
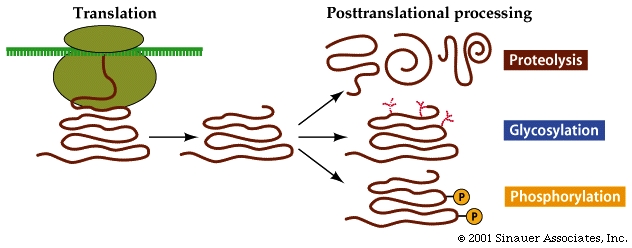
Some proteins require additional modification after synthesis before they become functional.
New chemical groups might be added to the protein, it might be folded (sometimes with he assistance of other proteins) or it might get trimmed.
As we have discussed previously, chemical signals in/on the proteins also, often direct them to their cellular destinations
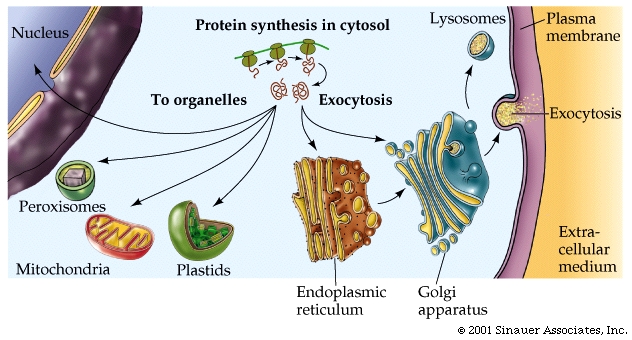
As the polypeptide chain forms, it spontaneously folds into its three-dimensional shape, mainly as a function of the various R groups that constitute the different amino acids within the polypeptide chain.
Those polypeptides that are destined to finish synthesis in the cytoplasm may contain information in their amino acid sequence that specifies where they belong: the nucleus, mitochondria, or peroxisomes.
Some of the proteins that are transported to a destination require chaperonin proteins and docking proteins at the membrane that the protein must interact with before it's ultimate destination can be assigned.
Those destined for the ER often generate an additional, hydrophobic "leader sequence" -approximately 25-amino acid long- that interacts with asignal recognition particle, which is composed of protein and RNA.
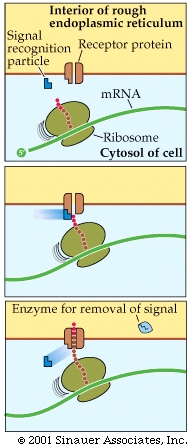
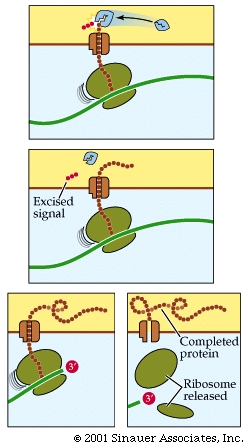
The association of the signal to the signal receptor particle stalls any additional translation until the immature protein that is still within the ribosome attaches to a specific receptor protein on the surface of the ER.
Translation continues with the protein moving through a pore in the ER membrane, thereafter the signal to go to the ER is often cleaved after the protein is extruded on to the interior of the ER (Endoplasmic reticulum).
In eukaryotes so many of the proteins that are synthesized are modified in some way AFTER translation. While these modifications ARE NOT unique to Eukaryotes, with eukaryotes it is the exception, not the rule when the finished, fully formed / folded protein is identical to the originally translated polypeptide product from the mRNA code.
Modification of proteins is often essential to the final functioning of the protein to "activate" its function.

Additional modifications, that are not specified by the synthesis process itself, also occur, and require additional changes to occur before the protein/enzyme can be "activated". The most obvious of these is "protease" dependent maturation, wherein part of the protein is cleaved to make a shortened, finished protein .
Insulin is an example of a protein that gets trimmed from proinsulin.

About Contact GSU Academics Degrees & Majors |
Admissions Undergraduate Research Dept. of Biology |
Libraries University Library Campus Life Housing |
Athletics Alumni
|

|